Welcome to the Ligandable Cysteine Database built with SLCABPP!
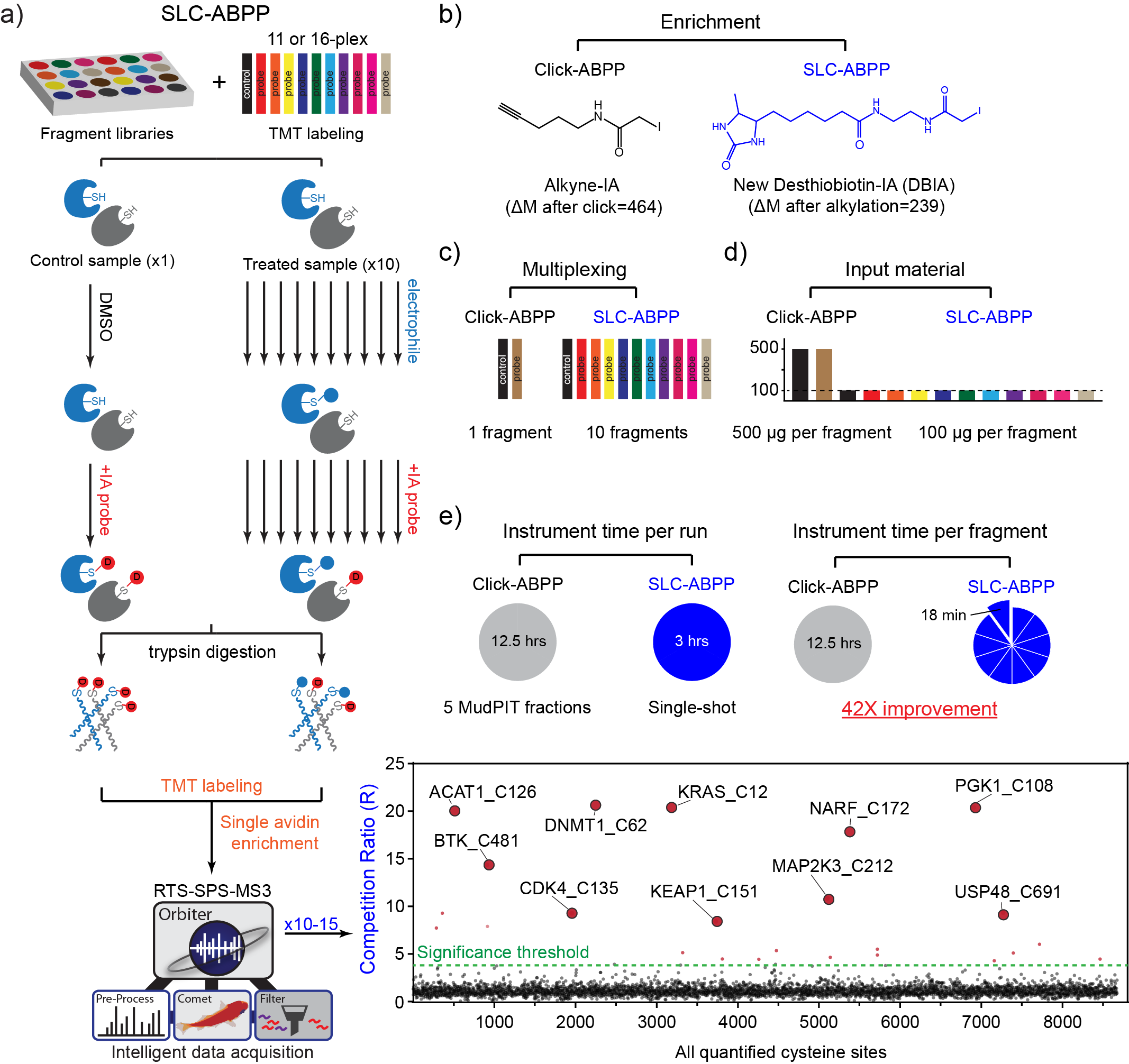
Current methods for measuring amino-acid side-chain reactivity lack the throughput needed to screen large chemical libraries for interactions across the proteome. Here we redesigned the workflow for activity-based protein profiling of reactive cysteine residues by using a smaller desthiobiotin-based probe, sample multiplexing, reduced protein starting amounts, and software to boost data acquisition in real-time on the mass spectrometer. Our method, Streamlined Cysteine Activity-Based Protein Profiling (SLC-ABPP), achieved a 42-fold improvement in sample throughput, corresponding to profiling library members at a depth of >8,000 reactive cysteine sites in 18 min per compound. We applied it to identify proteome-wide targets of covalent inhibitors to mutant KRASG12C and BTK. In addition, we created a resource of cysteine reactivity to 285 electrophiles in three human cell lines, which includes >20,000 cysteines from >6,000 proteins per line. The goal of proteome-wide profiling of cysteine reactivity across thousand-member libraries under several cellular contexts is now within reach.
Data Access and Information
Access data via the 'Data' tab above. Ligandable cysteines can be viewed from either a gene centric (select a gene of interest) or compound centric view (select a compound of interest).
Gene Centric
Selecting a gene of interest will populate the Site Map depicting the competition ratios of each site for each compound in this study. Clicking on the Site Map for a given compound will display the compounds structure and also populate the scatter plot at the bottom of the Data page to highlight all site competition ratios for that compound across the full dataset.
Compound Centric
Selecting a compound will populate the structure depecting and the plot at the bottom of the Data page to highlight all site competition ratios for that compound across the full dataset.
Citations
The SLCABPP web resource was built by Nathan Bulloch, David Nusinow, and Devin Schweppe
For more information see the publication: Kuljanin et al. Nat Biotech 2020
Or check out the Gygi Lab Website